There are multiple ways to categorize nearly everything in biology. The value of an individual categorization scheme depends on the perspective of the user. As Shirley Malcom (former Director, Education and Human Resources Programs, AAAS) once said, “We [Biologists] are splitters not lumpers.” This need for classification makes consensus difficult, can limit our way of thinking, and can make scientific findings sound more sensational than they really are. The increasing appreciation for the multifunctionality of biological entities at all scales reveals how difficult it is to put biology into exclusive and unambiguous categories. Furthermore, as multifunctionality becomes the norm, we should remember that it is the human in the role of researcher, clinician, scientific author, and reader of the scientific literature that needs classification to reveal the patterns and meaning in the complexity of living organisms. We should not let the current classification systems (ontologies) of genes, proteins, organelles, pathways, physiological systems, or even organisms limit how we think about and explore biology.
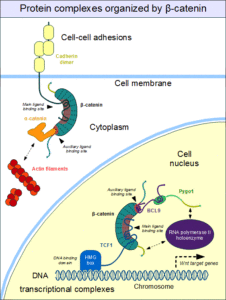
A few examples at each scale of biology illustrate the challenges created by our human predilection for classification. Just given the name of a protein or the abbreviation of the gene that encodes it, the first questions are likely to be: What is it? What does this protein do? If the name includes a function, then there is a reasonable chance of assuming the function implied by the name is correct and relevant. Otherwise, the gene abbreviation or protein name is unlikely to be meaningful unless it is very common in medicine, like insulin, or unless you have studied that gene or protein. Many proteins are multifunctional. The protein β-catenin is a good example. When incorporated into the protein complexes that allow cells to form stable contacts with each other, it is part of a cellular adhesion complex and so has the function of mediating cell-cell adhesion. In response to certain external signals, β-catenin can move into the nucleus and regulate gene expression. Thus, it is a transcriptional regulator. Certainly, β-catenin should be categorized with both functions. What if each function is important in a different context? Both functions need to be captured, but somehow the context-specific details need to be included as well.
The organization of proteins into distinct regulatory or biochemical pathways is also a human construction. Regulatory and signaling pathways are highly interconnected. Indeed, this must be true. Cells cannot move and divide at the same time, so the pathways controlling movement and division must be connected. Molecules previously considered as biochemical intermediates in metabolic pathways are becoming increasingly appreciated as regulators of signaling pathways and cellular behavior. How can this complexity in molecular function be captured in a useful way in a classification scheme?
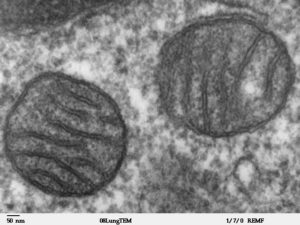
Moving up in size, the organelles in a eukaryotic cell tend to be functionally defined according to the function first identified or most studied. For example, textbooks describe mitochondria as the cell’s powerhouse, because mitochondria generate ATP; but the mitochondria are a source of reactive oxygen species and many kinds of intracellular signaling molecules and are a sink for calcium. So, what is the best way to classify mitochondrial function?
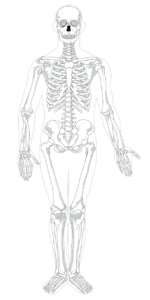
Moving even farther up in size, organs are classified into physiological systems—the cardiovascular system, the endocrine system, the musculoskeletal system and so on. An excellent example is bone. As the skeletal system, bones provide support, movement, and protection. However, bones are also part of the immune system: Bones are the site of blood cell production. Bones are part of the endocrine system: They release hormones that regulate appetite, fertility, and metabolism. Even the well-known and long-standing physiological categories fail to represent a true picture of the complex multifunctionality of the tissues and cells that comprise organ systems.
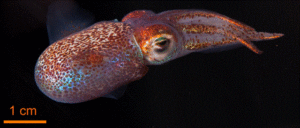
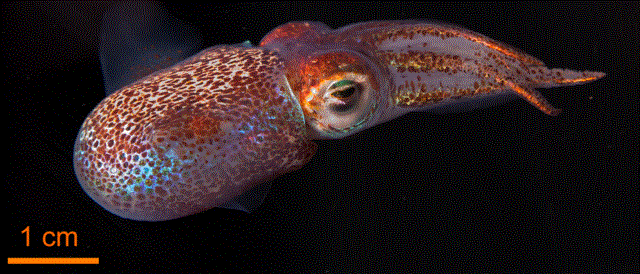
Going all the way to a person, plants, and marine organisms, these are defined by a single species name; yet people have microbiomes in their gut, mouth, skin, ears, eyes, and genitals; legumes have symbiotic fungi that are part of their root systems; and many bioluminescent marine animals have bacteria that provide the light. So, how should we classify these? They, indeed even we humans, are all metaorganisms—multiple species living in harmony.
Why does it matter if it is hard to classify biological information? Classification enables systems-level analysis of large data sets. Classification enables automation. Classification increases the ability to retrieve information from large data sets and enables the interpretation, discovery of new patterns, and acquisition of knowledge from large data sets. However, information acquired through use of classification schemes is only as good as the classification scheme, the consistency with which it is applied, and knowledge about its limitations.
Ideally, all functionally important information should be included whenever possible in the scientific literature. Furthermore, the relevant context-specific function(s) should be indicated when known. This need for context-specific information to ensure accuracy means that using text-mining and then applying an ontology that includes all functional classifications is not going to provide the necessary context-specific information. Automated classification is challenging and curation is necessary to ensure context-dependent accuracy. Thus, effective scientific communication relies on the author to provide the contextual details to ensure that the literature is accurate and precise, which makes biological findings as reproducible as possible.
Cite as: N. R. Gough, Challenges in Classifying Biology. BioSerendipity (5 August 2017). https://www.bioserendipity.com/challenges-in-classifying-biology/